Reorganizing the RNA Polymerase II Complex for Replication of an Infectious Noncoding RNA in Vivo: Key Findings Explained
What if a pathogen could repurpose your body’s core genetic machinery to copy its own genome — even when that genome produces no proteins? This is the reality of infectious noncoding RNA (ncRNA) pathogens, which have evolved a clever strategy to hijack the RNA polymerase II (Pol II) complex for replication in vivo. New research into how these pathogens reorganize Pol II is reshaping our understanding of viral-host interactions and opening doors to novel antiviral therapies.
What Is the RNA Polymerase II Complex?
RNA polymerase II is a multi-subunit enzyme found in the nucleus of all eukaryotic cells, responsible for transcribing protein-coding genes into messenger RNA (mRNA). A typical Pol II complex contains 12 core subunits, plus additional regulatory factors that control its activity.
Normally, Pol II uses DNA as a template — it cannot copy RNA genomes on its own. It also primarily produces mRNA that is translated into proteins, not standalone noncoding RNA. This makes its role in ncRNA replication all the more surprising.
Infectious Noncoding RNA: A Unique Pathogen Class
Infectious ncRNAs include agents like viroids (small plant pathogens) and hepatitis delta virus (HDV, a human pathogen that co-infects with hepatitis B). Unlike most RNA viruses, these pathogens do not encode their own RNA-dependent RNA polymerase (RdRP) to copy their genomes.
Instead, they rely entirely on host cellular machinery to replicate. For years, researchers were puzzled by how these RNA-only pathogens could copy themselves without a dedicated viral polymerase — until recent work revealed the critical role of Pol II reorganization.
Reorganizing the Pol II Complex for In Vivo ncRNA Replication
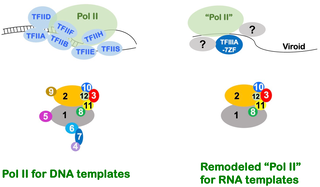
The core finding of recent studies is that infectious ncRNAs trigger specific, reversible changes to the Pol II complex that allow it to use RNA as a template instead of DNA. These changes are tailored to support replication in living organisms (in vivo), not just artificial lab conditions.
Key Structural Changes Driving Reorganization
- Subunit swapping: Non-essential Pol II subunits are replaced with host or viral factors that enable the complex to bind RNA templates directly.
- CTD phosphorylation shifts: The C-terminal domain (CTD) of Pol II’s largest subunit undergoes specific phosphorylation changes that switch the complex from transcription mode to RNA replication mode.
- Nuclear compartmentalization: Reorganized Pol II complexes relocate to specialized nuclear speckles or virolipid droplets where ncRNA replication is concentrated, avoiding interference with normal host transcription.
- Template recognition adjustments: Associated factors modify the complex’s active site to accommodate the unique secondary structure of infectious ncRNAs.
Critically, these changes are transient and specific to infected cells — healthy cells maintain normal Pol II function for transcription, minimizing collateral damage to host gene expression.
Why This Research Matters for Global Health
Hepatitis delta virus alone affects an estimated 15–20 million people worldwide, causing the most severe form of viral hepatitis with no curative treatment available. Current therapies target hepatitis B, but HDV persists even after HBV is suppressed.
By targeting the specific Pol II reorganization steps required for HDV replication, researchers could develop drugs that block ncRNA copying without disrupting normal host transcription. This approach would also apply to other infectious ncRNA pathogens, including emerging plant viroids that threaten global food security.
Current Gaps and Future Research Directions
Most of our current understanding comes from cell culture and model organism studies — more work is needed to confirm these mechanisms in human patients. Researchers are also working to map the exact viral and host factors that trigger Pol II reorganization, to identify druggable targets.
Future studies will also explore whether this reorganization mechanism is used by other noncoding pathogens, including some emerging zoonotic viruses with noncoding RNA genomes.
Reorganizing the RNA polymerase II complex represents a remarkable example of pathogen adaptation, turning a core host function against the cell itself. As we uncover more details of this process in vivo, we move closer to new treatments for some of the world’s most persistent infectious diseases. For now, this work reminds us that even the most well-studied cellular machinery can hold surprising secrets — especially when challenged by infectious noncoding RNAs.

Comments are closed, but trackbacks and pingbacks are open.